Integrative Omics in Parkinson Disease

Team: Joanne Trinh, PhD (Group leader); Theresa Lüth (PhD candidate); Carolin Gabbert (PhD candidate); Susen Schaake (lab manager, research technician); Christoph Much (Research technician); Joshua Laß (PhD candidate); André Fienemann (PhD candidate); Natascha Bahr (Documentalist); Yuliia Kanana (Technician); Julia Charlotte Prietzsche (Master’s student).
The team’s main interest is the identification of novel causal genes, genetic risk factors and penetrance modifiers using family studies, association and epidemiology with integrative large-scale omics approaches.
Novel genes and structural variants implicated in Parkinson’s disease (PD) and disease-associated pathways
In an effort to improve understanding of PD and impact of patient care, we initiated an early onset PD study to identify novel genes and cellular pathways associated with this disease. After an initial exome sequencing study, the Oxford Nanopore third generation sequencing technology will be used to elucidate the ‘dark matter’ of the human genome to identify novel candidate PD genes in inherited (familial) forms and in patients with sporadic PD.
Genetic modifiers of monogenic forms of Parkinson’s disease
Our interest lies in finding genetic modifiers that explain the clinical variability of LRRK2, Parkin and PINK1 forms of PD. We use a series of integrative omics approaches (genome/exome sequencing, deep mitochondrial sequencing, RNA sequencing and epigenetic markers) to further elucidate genetic modifiers or disease. The identification of functional pathogenic variants may also direct prevention strategies and nominate molecular targets for individualized disease-modifying therapies.
Genetic and environmental LRRK2 age-at-onset modifiers
Understanding factors and underlying mechanisms that influence reduced penetrance/expressivity has a high imperative given that these discoveries provide the rationale, molecular insight and research tools to develop neuroprotective and disease-modifying therapies. Linkage analysis has been previously performed on age at onset for Tunisian Arab Berber LRRK2 families. We hypothesize that 1) there are variants in candidate genes within unexplored linkage regions that influence age at onset 2) candidate genes identified will replicate in a larger LRRK2 cohort 3) the specific variants of interest influence transcriptional regulation and expression of gene candidates.
Furthermore, we are investigating the environmental and lifestyle factors that are related to LRRK2. Geoparkinson Questionnaire and PD Risk Factor Questionnaire (RFQ-U) is used to assess environmental and lifestyle factors. Motor and non-motor symptoms is evaluated by the Movement Disorder Society Unified Parkinson’s Disease Rating Scale (MDS UPDRS), Hoehn and Yahr, and the Scales for Outcomes in Parkinson’s Disease – Autonomic Dysfunction (SCOPA-AUT) scores. Our data suggests that tobacco use and black tea consumption have a protective effect on the age-dependent penetrance in LRRK2 parkinsonism. Moreover, tobacco users show differences in depressed, anxious mood and apathy.
The role of the extranuclear DNA genome of the mitochondrion
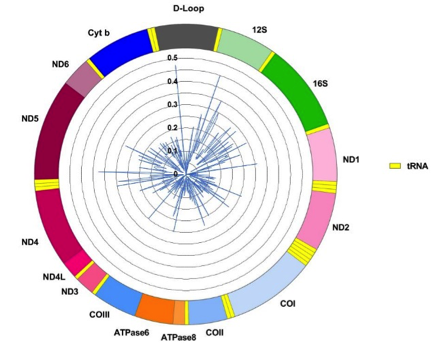
In collaboration with researchers at the University of Luxembourg (Prof. Anne Grünewald, LCSB), we use deep sequencing and third generation sequencing of the mtDNA to measure mtDNA integrity (variants and modifications) in monogenic PD (Parkin, PINK1), idiopathic PD, in-vitro models of PD (IPSC-derived neurons) and other parkinsonism-related syndromes such as POLG and Twinkle pathogenic mutation carriers.
Integrative modelling of the microbiome and mitochondria to detect progression markers for Parkinson’s disease
Monogenic forms of PD and strong risk factors account for ~10% of patients worldwide. Common variants in PD and mitochondrial genes constitute polygenic risk scores (PRS), which can predict an up to 6-fold increased risk of iPD. Among others, monogenic forms are caused by mutations in LRRK2, PINK1 and Parkin, which mediate a variety of biological functions involving mitochondria. PRS in iPD, in addition to homogeneous monogenic forms, allow patient stratification to identify and characterize progression markers. Similar to mtDNA, the microbiome is inherited from the mother, but then develops into a unique signature. There is bidirectional communication between the intestinal microbiome and the mitochondria through endocrine, immunological or humoral pathways. The idea that microbiota are associated with neurological disorders is well established, but how the microbiome develops and influences the progression of PD is unknown and can only be elucidated in longitudinal studies. We have initiated a metagenomic and metranscriptomic study on PD progression.
We value and acknowledge scientific collaborations from Prof. Dr. Matthew Farrer, Prof. Dr. Anne Grünewald, Prof. Dr. Andrew Hicks, Prof. Dr. Peter Pramstaller, Prof. Dr. Inke König, Dr. Patrick May, Prof. Bastiaan Bloem, Prof. Dr. Hauke Busch, Prof. Dr. Jan Aasly, Prof. Dr. Jan Rupp, Dr. Simon Graspeuntner
- Translational Neurogenetics
- Genetics of Rare Diseases
- Functional Genetics of Movement Disorders
- Integrative Omics in Parkinson Disease
- Neuropsychiatric Epidemiology
- Clinical Neurogenetics and Neuroimaging
- Mitochondrial function in movement disorders
- Institut für Systemische Motorikforschung
- Applied Stem Cell Biology